16.15: Problems for Chapter 16
- Page ID
- 2453
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\( \newcommand{\dsum}{\displaystyle\sum\limits} \)
\( \newcommand{\dint}{\displaystyle\int\limits} \)
\( \newcommand{\dlim}{\displaystyle\lim\limits} \)
\( \newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\)
( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\id}{\mathrm{id}}\)
\( \newcommand{\Span}{\mathrm{span}}\)
\( \newcommand{\kernel}{\mathrm{null}\,}\)
\( \newcommand{\range}{\mathrm{range}\,}\)
\( \newcommand{\RealPart}{\mathrm{Re}}\)
\( \newcommand{\ImaginaryPart}{\mathrm{Im}}\)
\( \newcommand{\Argument}{\mathrm{Arg}}\)
\( \newcommand{\norm}[1]{\| #1 \|}\)
\( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\)
\( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\AA}{\unicode[.8,0]{x212B}}\)
\( \newcommand{\vectorA}[1]{\vec{#1}} % arrow\)
\( \newcommand{\vectorAt}[1]{\vec{\text{#1}}} % arrow\)
\( \newcommand{\vectorB}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\( \newcommand{\vectorC}[1]{\textbf{#1}} \)
\( \newcommand{\vectorD}[1]{\overrightarrow{#1}} \)
\( \newcommand{\vectorDt}[1]{\overrightarrow{\text{#1}}} \)
\( \newcommand{\vectE}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash{\mathbf {#1}}}} \)
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \)
\(\newcommand{\longvect}{\overrightarrow}\)
\( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)
\(\newcommand{\avec}{\mathbf a}\) \(\newcommand{\bvec}{\mathbf b}\) \(\newcommand{\cvec}{\mathbf c}\) \(\newcommand{\dvec}{\mathbf d}\) \(\newcommand{\dtil}{\widetilde{\mathbf d}}\) \(\newcommand{\evec}{\mathbf e}\) \(\newcommand{\fvec}{\mathbf f}\) \(\newcommand{\nvec}{\mathbf n}\) \(\newcommand{\pvec}{\mathbf p}\) \(\newcommand{\qvec}{\mathbf q}\) \(\newcommand{\svec}{\mathbf s}\) \(\newcommand{\tvec}{\mathbf t}\) \(\newcommand{\uvec}{\mathbf u}\) \(\newcommand{\vvec}{\mathbf v}\) \(\newcommand{\wvec}{\mathbf w}\) \(\newcommand{\xvec}{\mathbf x}\) \(\newcommand{\yvec}{\mathbf y}\) \(\newcommand{\zvec}{\mathbf z}\) \(\newcommand{\rvec}{\mathbf r}\) \(\newcommand{\mvec}{\mathbf m}\) \(\newcommand{\zerovec}{\mathbf 0}\) \(\newcommand{\onevec}{\mathbf 1}\) \(\newcommand{\real}{\mathbb R}\) \(\newcommand{\twovec}[2]{\left[\begin{array}{r}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\ctwovec}[2]{\left[\begin{array}{c}#1 \\ #2 \end{array}\right]}\) \(\newcommand{\threevec}[3]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\cthreevec}[3]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \end{array}\right]}\) \(\newcommand{\fourvec}[4]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\cfourvec}[4]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \end{array}\right]}\) \(\newcommand{\fivevec}[5]{\left[\begin{array}{r}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\cfivevec}[5]{\left[\begin{array}{c}#1 \\ #2 \\ #3 \\ #4 \\ #5 \\ \end{array}\right]}\) \(\newcommand{\mattwo}[4]{\left[\begin{array}{rr}#1 \amp #2 \\ #3 \amp #4 \\ \end{array}\right]}\) \(\newcommand{\laspan}[1]{\text{Span}\{#1\}}\) \(\newcommand{\bcal}{\cal B}\) \(\newcommand{\ccal}{\cal C}\) \(\newcommand{\scal}{\cal S}\) \(\newcommand{\wcal}{\cal W}\) \(\newcommand{\ecal}{\cal E}\) \(\newcommand{\coords}[2]{\left\{#1\right\}_{#2}}\) \(\newcommand{\gray}[1]{\color{gray}{#1}}\) \(\newcommand{\lgray}[1]{\color{lightgray}{#1}}\) \(\newcommand{\rank}{\operatorname{rank}}\) \(\newcommand{\row}{\text{Row}}\) \(\newcommand{\col}{\text{Col}}\) \(\renewcommand{\row}{\text{Row}}\) \(\newcommand{\nul}{\text{Nul}}\) \(\newcommand{\var}{\text{Var}}\) \(\newcommand{\corr}{\text{corr}}\) \(\newcommand{\len}[1]{\left|#1\right|}\) \(\newcommand{\bbar}{\overline{\bvec}}\) \(\newcommand{\bhat}{\widehat{\bvec}}\) \(\newcommand{\bperp}{\bvec^\perp}\) \(\newcommand{\xhat}{\widehat{\xvec}}\) \(\newcommand{\vhat}{\widehat{\vvec}}\) \(\newcommand{\uhat}{\widehat{\uvec}}\) \(\newcommand{\what}{\widehat{\wvec}}\) \(\newcommand{\Sighat}{\widehat{\Sigma}}\) \(\newcommand{\lt}{<}\) \(\newcommand{\gt}{>}\) \(\newcommand{\amp}{&}\) \(\definecolor{fillinmathshade}{gray}{0.9}\)Link to Solution Manual
P16.1: During intense exercise, pyruvate is converted to lactate in muscle tissue. It is the lactate that you can blame for the sore muscles you feel the day after a workout.
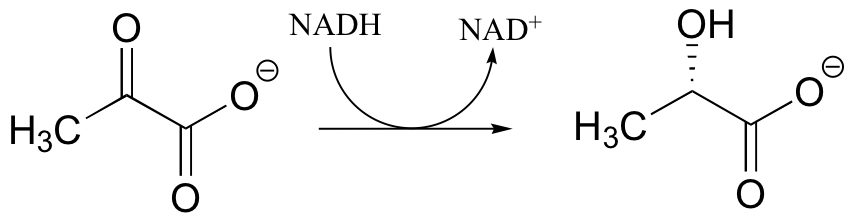
a) Draw arrows showing the electron movement in this reaction, and include the electron movement in the nicotinamide ring of the NADH coenzyme.
b) Which face of pyruvate is attacked by the hydride from NADH?
P16.2: Show a mechanism for the redox reactions below. In part a, include the electron movement in the nicotinamide ring of the coenzyme.
a)
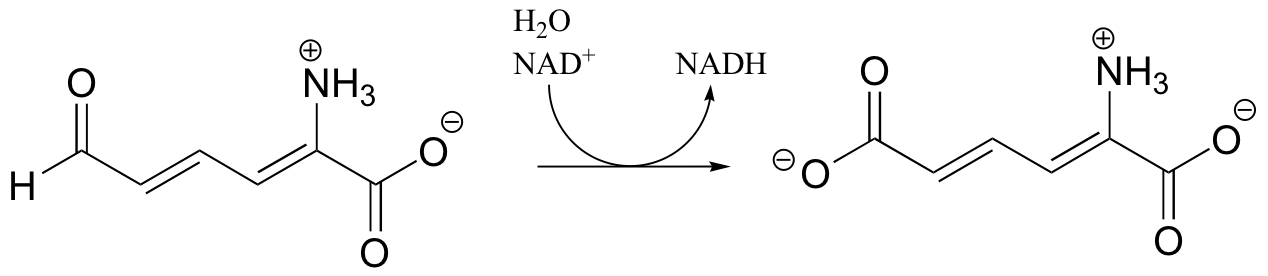
b)
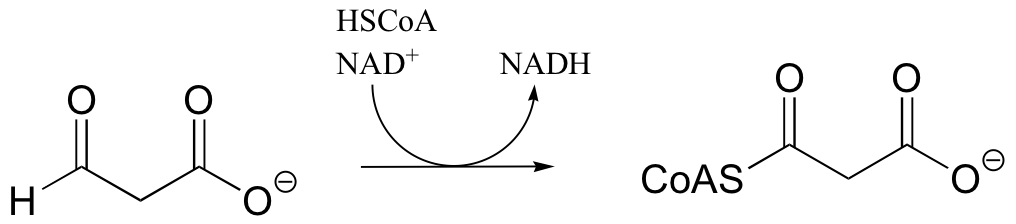
P16.3: In the transformation below, the side chain of aspartate is altered, but the main peptide chain is not affected. Show the most probable structure of species A and B.
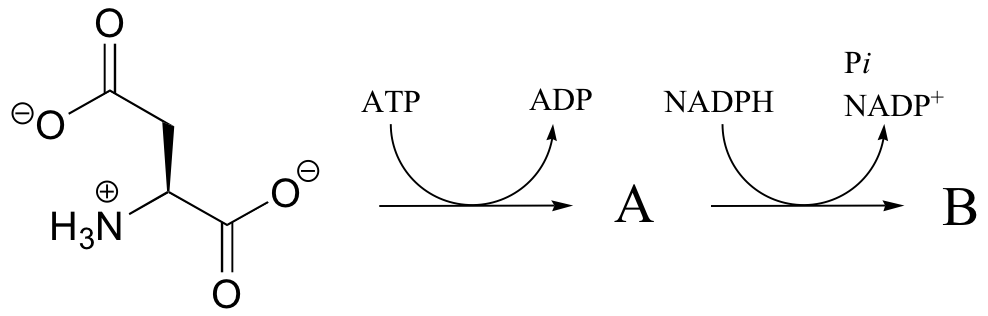
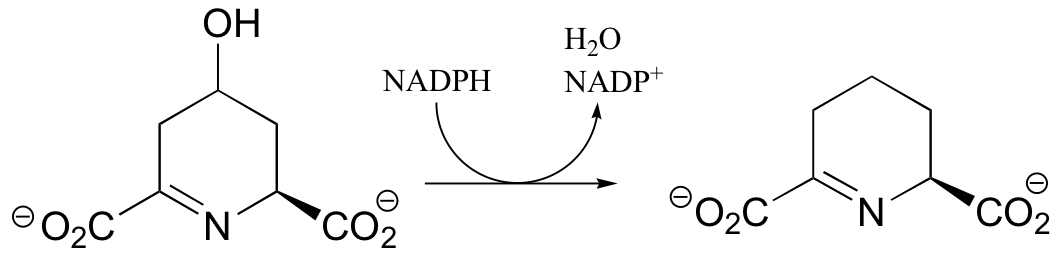
P16.4: Propose mechanisms for the following transformations:
a) From lysine biosynthesis:
b) This is a PLP-dependent step in the glycine degradation pathway. Start with the glycine-PLP adduct, and end with the enzyme-PLP adduct.
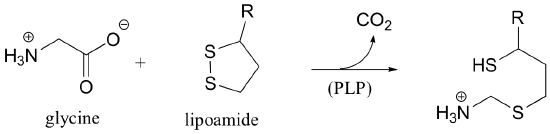
P16.5: The following is a reaction in RNA and DNA biosynthesis. Propose a mechanism, including the structure of and role played by the redox coenzyme.
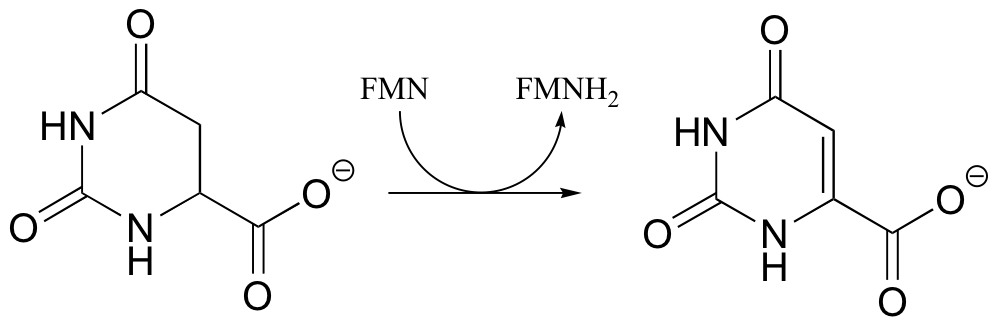
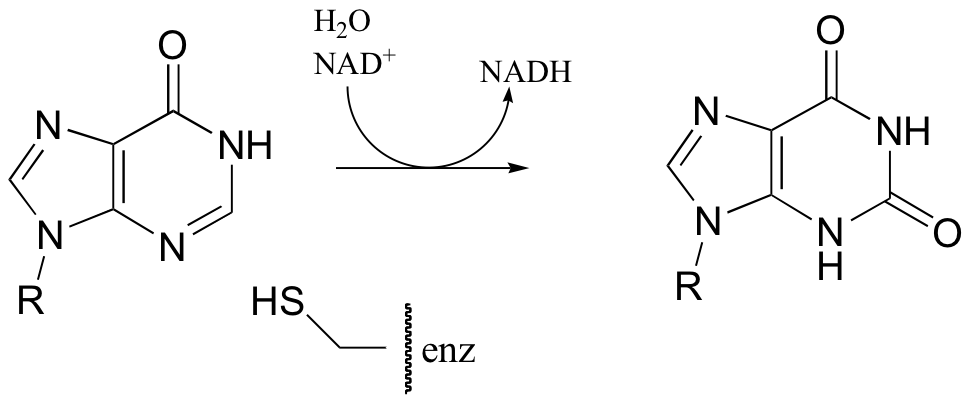
P16.6: The following reaction in the guanosine ribonucleotide biosynthetic pathway involves a covalent cysteine-linked enzyme-substrate intermediate. Propose a mechanism.
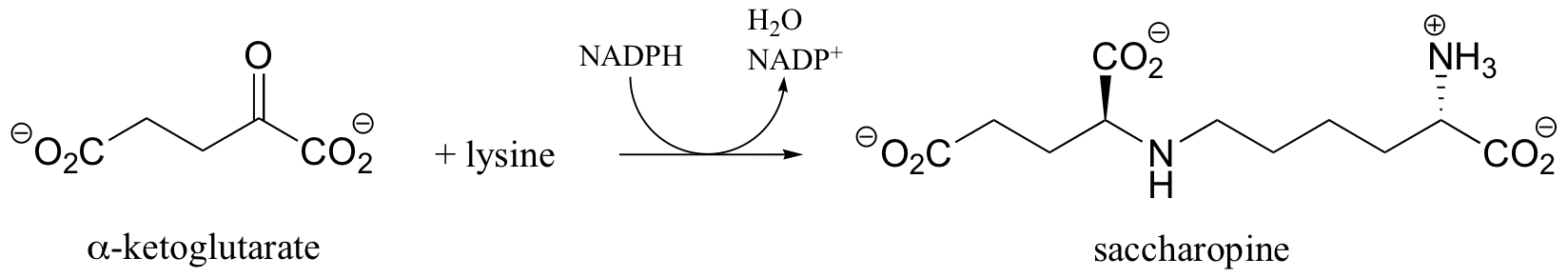
P16.7: The first step in the lysine degradation pathway is a reductive condensation with a-ketoglutarate to form an intermediate called saccharopine.
a) Propose a mechanism for this transformation.
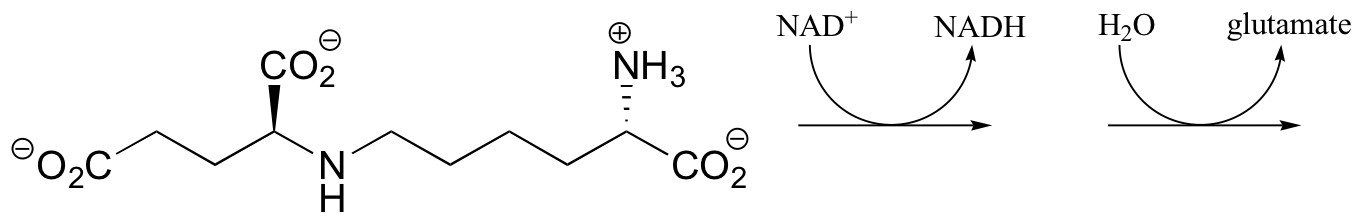
b) Saccharopine (see the structure in part a) of this problem) is broken up to yield glutamate and a second product that contains an aldehyde group. Predict the structure of this second product, and show a mechanism for the reaction.
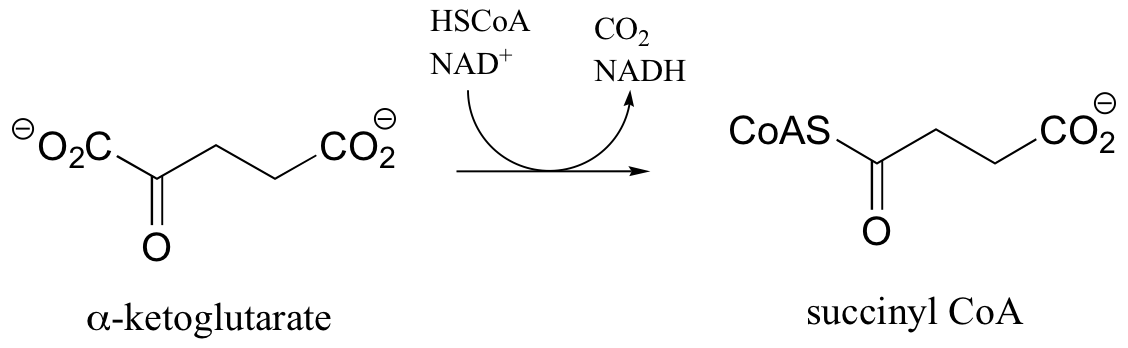
P16.8: The reaction below, from the citric acid cycle, is analogous to a very important reaction that we have already studied. Identify that reaction, as well as three cofactors that participate in the transformation but are not specified below (they are not shown because they are not used up, but rather are regenerated as part of the reaction’s catalytic cycle).
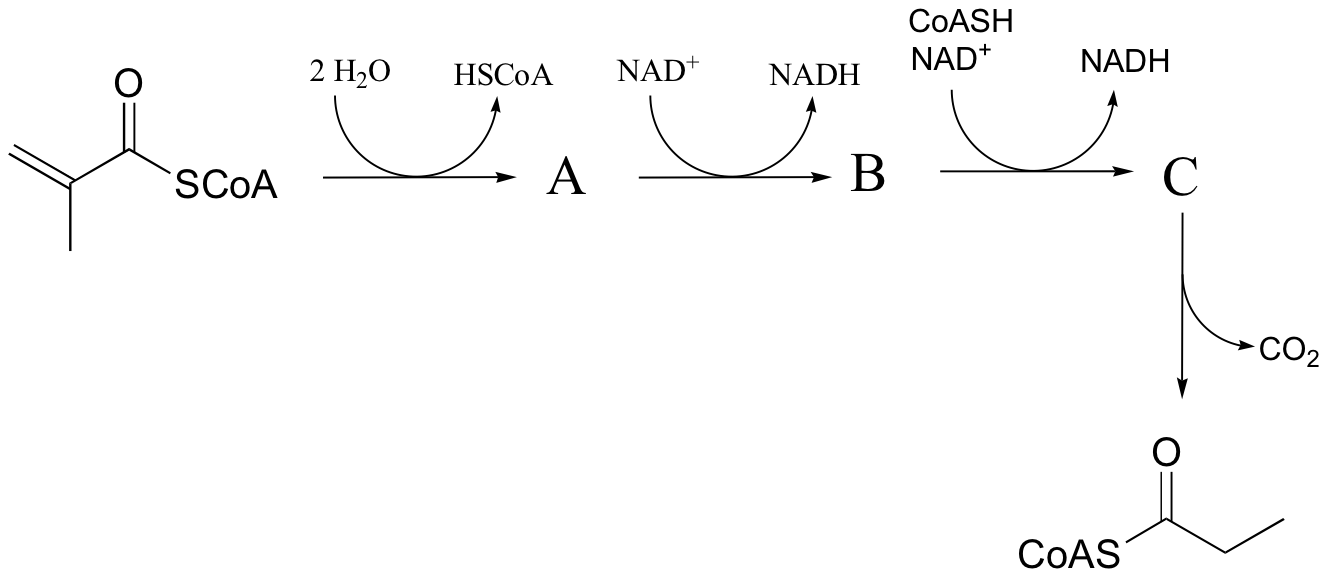
P16.9: Predict the structures of species A and B in the pathway below.
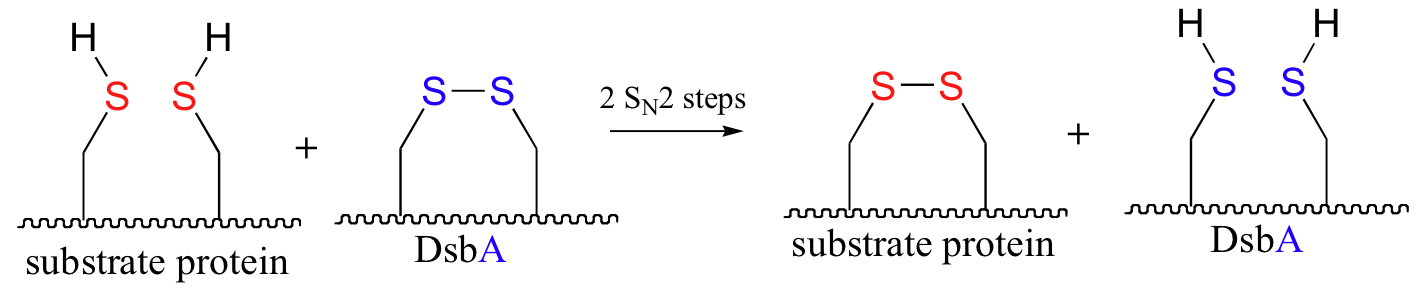
P16.10: An enzyme called DsbA is responsible for the formation of disulfide bonds in bacterial proteins. The process - which can be thought of as a 'disulfide exchange', involves the cleavage of a disulfide bond between two active site cysteines in DsbA. It is accomplished through two successive SN2 displacements.
DsbA is then returned to it's starting (disulfide) state through a second disulfide exchange reaction with another protein called DsbB.
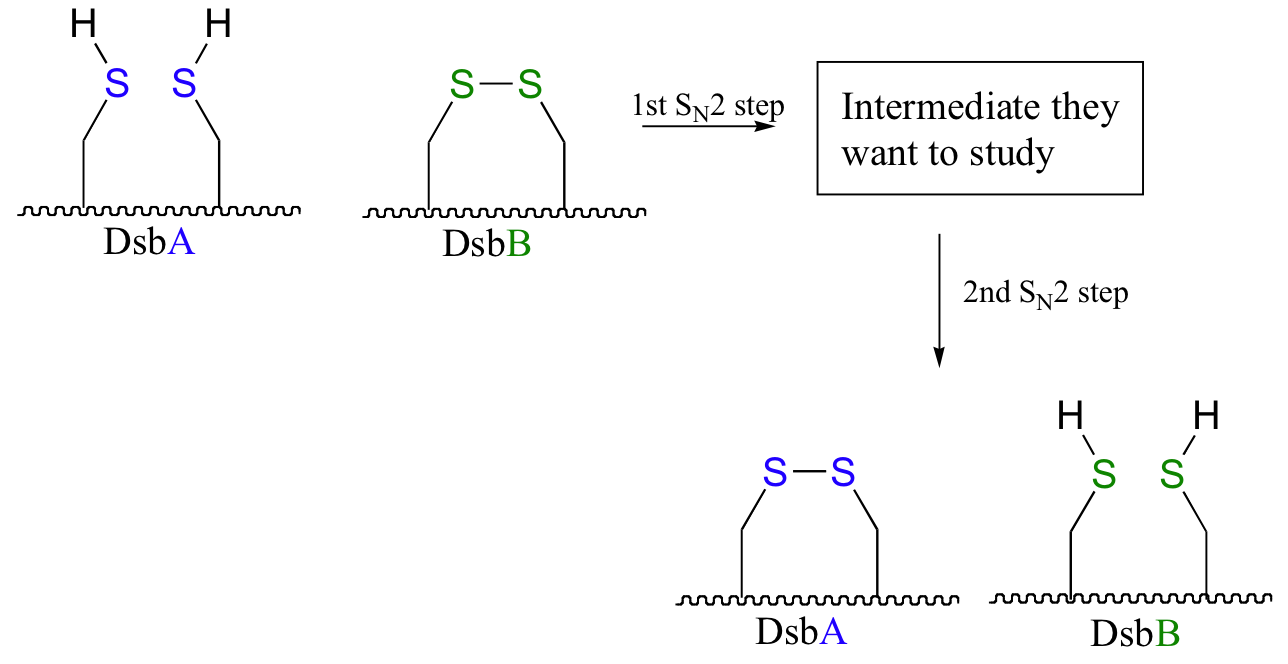
Scientists were interested in studying the intermediate species in this reaction, but found that it is very short-lived and difficult to isolate. In order to address this problem, they ran the reaction with a synthetic analog of DsbB that contained an unnatural bromoalanine amino acid in place of one of the active site cysteines.
a) Draw a complete mechanism for the disulfide exchange reaction between DsbA and DsbB.
b) Show how the bromoalanine-containing DsbB analog allowed for the isolation of an intermediate that resembles the true, short-lived intermediate.
P16.11: In section 15.6B we learned about ortho/para- vs meta-directing groups in electrophilic aromatic substitution reactions. Discuss the regiochemistry of the kynurenine hydroxylation reaction (section 16.10A) in this context.
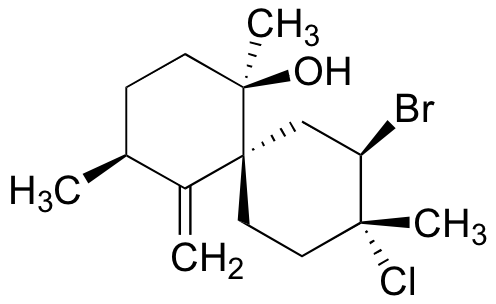
P16.12: Show a possible starting material and mechanism for the enzymatic halogenation reaction that results in the formation of the natural product below, which was isolated from marine red algae. Assume that the first step is the oxidation of Br- to Br+.
P16.13: In section 17.2E we will learn how ascorbic acid (vitamin C) acts as a 'radical scavenger' antioxidant to protect our cells from damage by free radical species. In doing so, it is converted to dehydroascorbate . One possible metabolic fate of dehydroascorbate is to be recycled back to ascorbic acid by the action of glutathione - this can happen either through and enzyme catalyzed reaction or through an enzyme-free reaction between glutathione and dehydroascorbate.
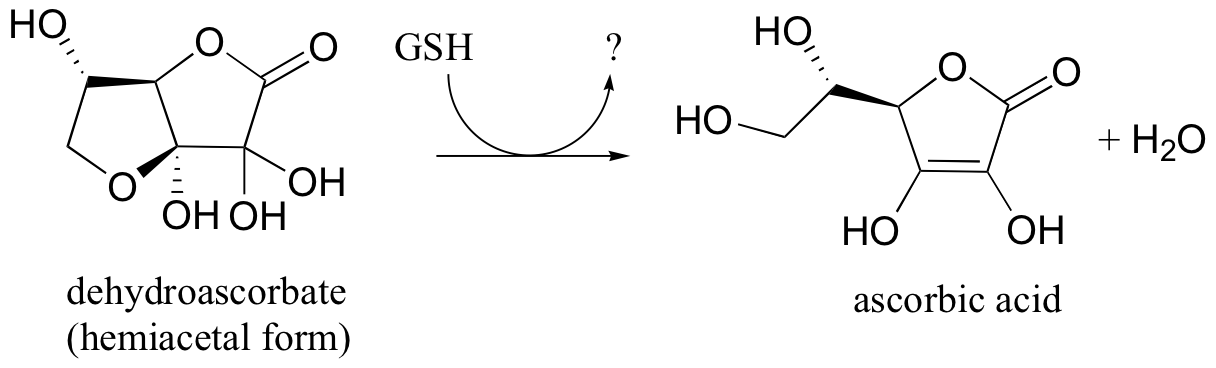
Suggest a likely mechanism for the enzyme-free reaction.
P16.14: Predict the products of the following laboratory reactions:
a)
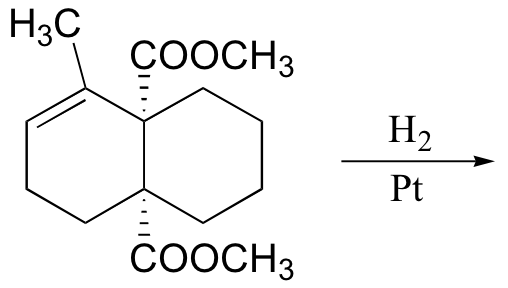
b)
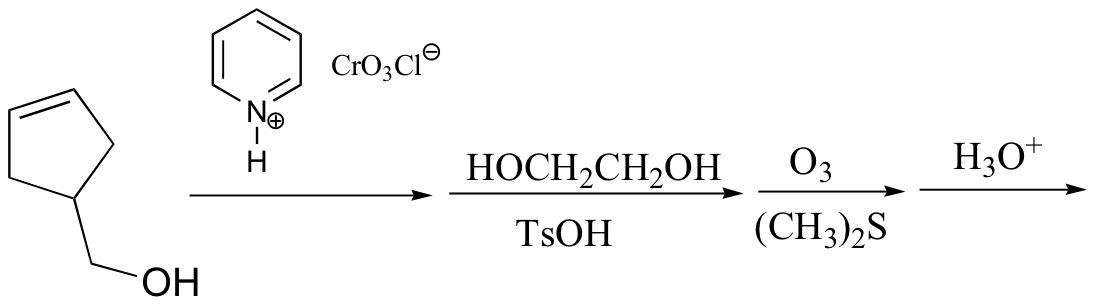
c) (give the structures of compounds A and B)
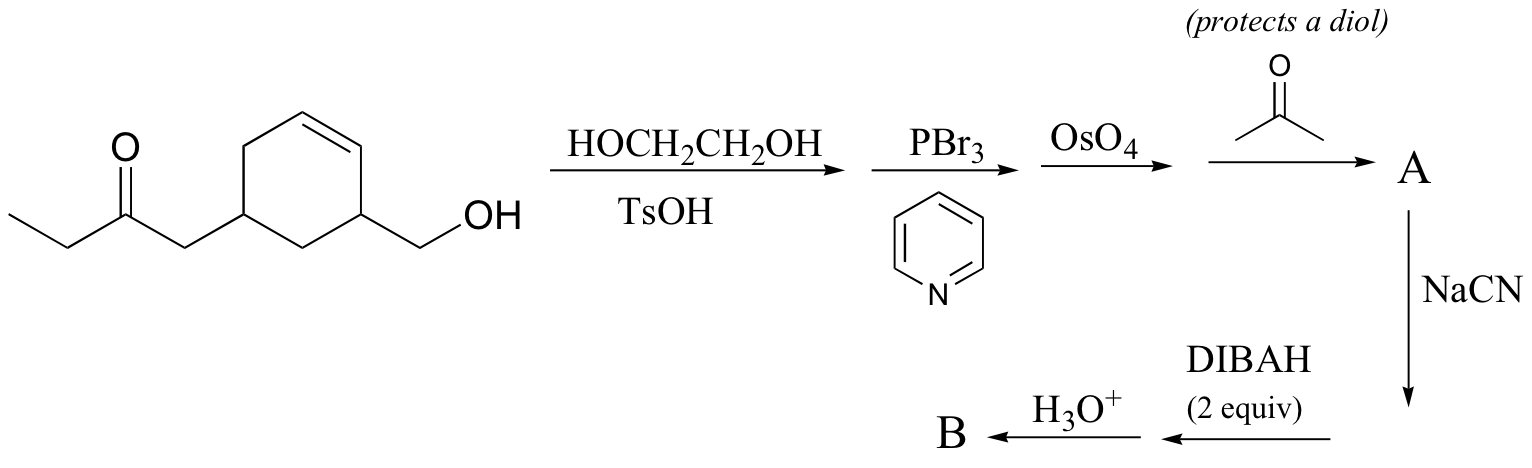
d) (give the structures of A, B, and C)
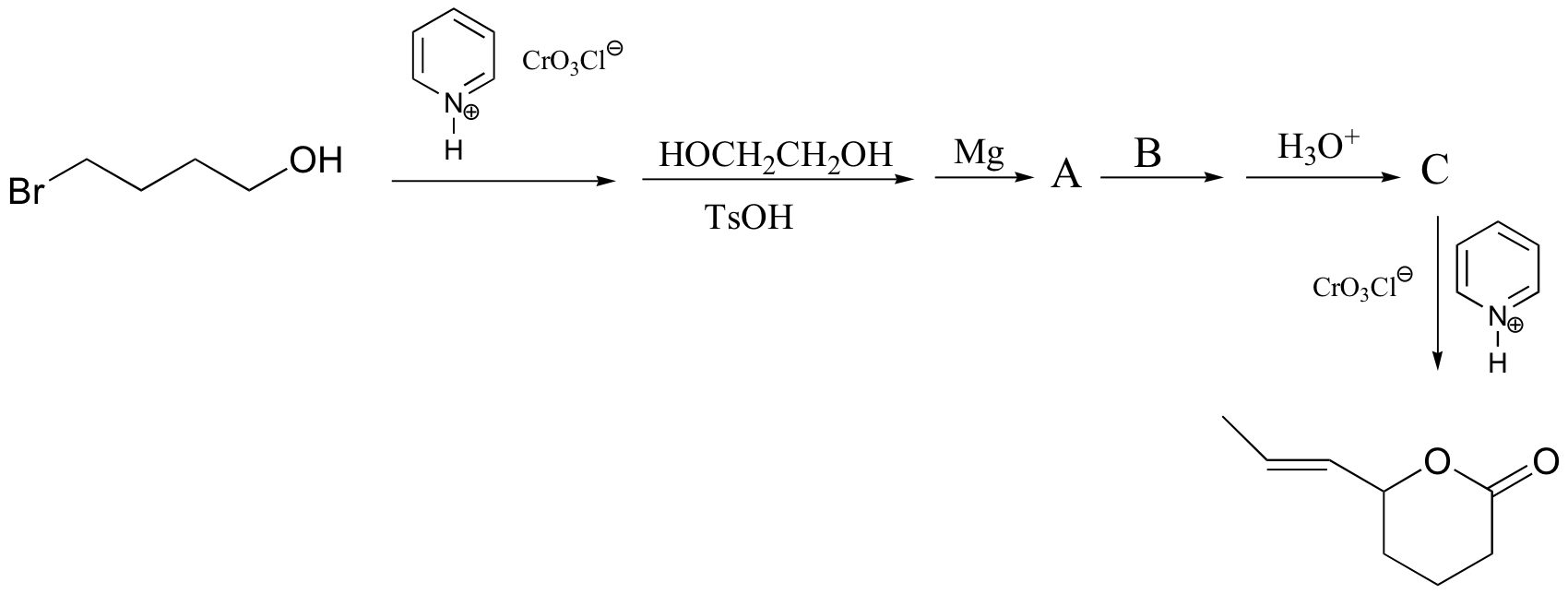
P16.15: Propose routes for the following multistep syntheses. You may use any synthetic reagents covered in the text so far, along with any organic starting materials in addition to those given in the problems.
a) (hint - remember, Friedel-Crafts alkylation reactions generally don't work well with primary alkyl halides, as they undergo rearrangements. Try an acylation/reduction combination. Think carefully about ring directing effects when deciding the order of your reactions!)
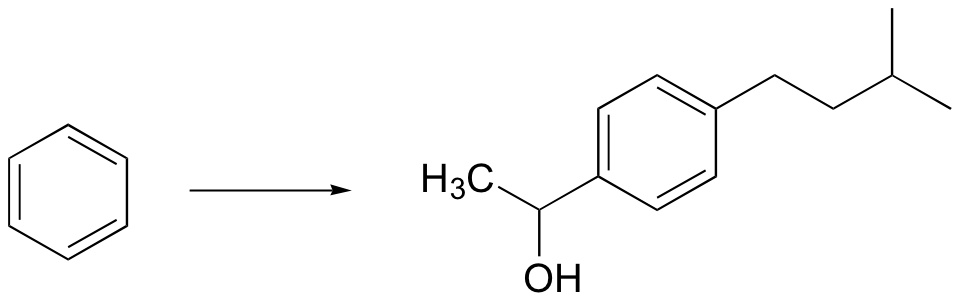
b)
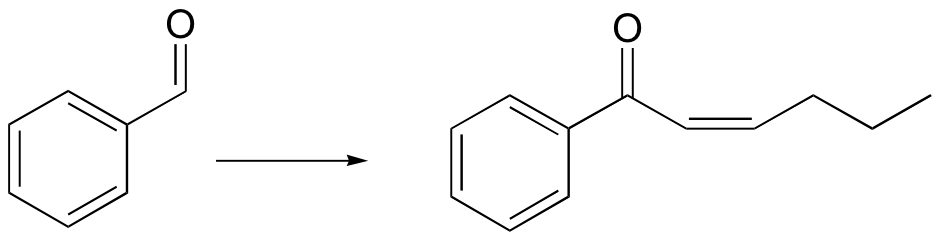
c)
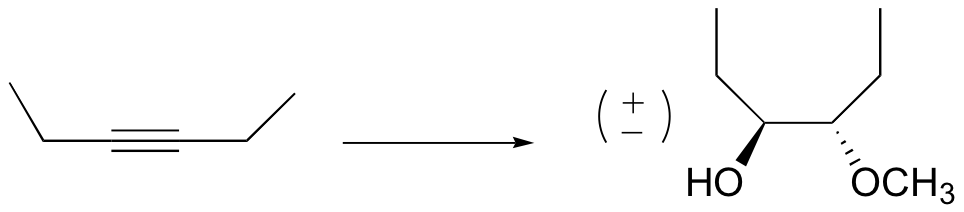
P16.16: Propose a mechanism for a) the Swern oxidation, and b) the PCC oxidation of 1-butanol.
Challenge problems
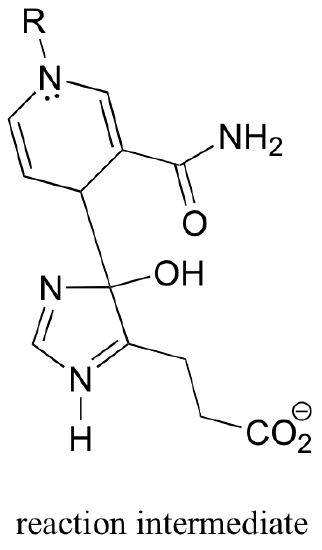
C16.1: As we have seen in this chapter, the biochemical role of NAD+ is to act as a hydride acceptor in dehydrogenation reactions. An exception is the reaction catalyzed by the histidine degradation pathway enzyme urocanase.
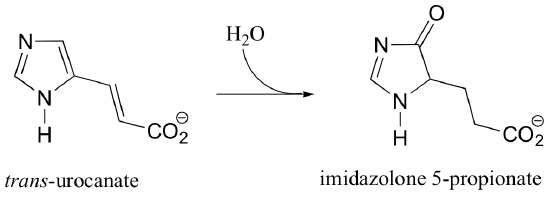
In this reaction, NAD+ acts as a catalytic, electron-sink coenzyme - it temporarily accepts electrons from a pi bond in the substrate, resulting in a covalent substrate-NAD adduct. This allows a key isomerization step to occur on the substrate through a protonation-deprotonation mechanism, followed by addition of water, cleavage of the substrate-NAD adduct to regenerate NAD+, and finally tautomerization to the product. Propose a mechanism that fits this description, and involves the intermediate below.