13.1: Error de muestreo (Sección @ref {samplingerror})
- Page ID
- 150697
\( \newcommand{\vecs}[1]{\overset { \scriptstyle \rightharpoonup} {\mathbf{#1}} } \) \( \newcommand{\vecd}[1]{\overset{-\!-\!\rightharpoonup}{\vphantom{a}\smash {#1}}} \)\(\newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\) \( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\) \( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\) \( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\) \( \newcommand{\Span}{\mathrm{span}}\) \(\newcommand{\id}{\mathrm{id}}\) \( \newcommand{\Span}{\mathrm{span}}\) \( \newcommand{\kernel}{\mathrm{null}\,}\) \( \newcommand{\range}{\mathrm{range}\,}\) \( \newcommand{\RealPart}{\mathrm{Re}}\) \( \newcommand{\ImaginaryPart}{\mathrm{Im}}\) \( \newcommand{\Argument}{\mathrm{Arg}}\) \( \newcommand{\norm}[1]{\| #1 \|}\) \( \newcommand{\inner}[2]{\langle #1, #2 \rangle}\) \( \newcommand{\Span}{\mathrm{span}}\)\(\newcommand{\AA}{\unicode[.8,0]{x212B}}\)
Aquí vamos a muestrear repetidamente de la variable Altura NHANES para obtener la distribución muestral de la media.
sampSize <- 50 # size of sample
nsamps <- 5000 # number of samples we will take
# set up variable to store all of the results
sampMeans <- tibble(meanHeight=rep(NA,nsamps))
# Loop through and repeatedly sample and compute the mean
for (i in 1:nsamps) {
sampMeans$meanHeight[i] <- NHANES_adult %>%
sample_n(sampSize) %>%
summarize(meanHeight=mean(Height)) %>%
pull(meanHeight)
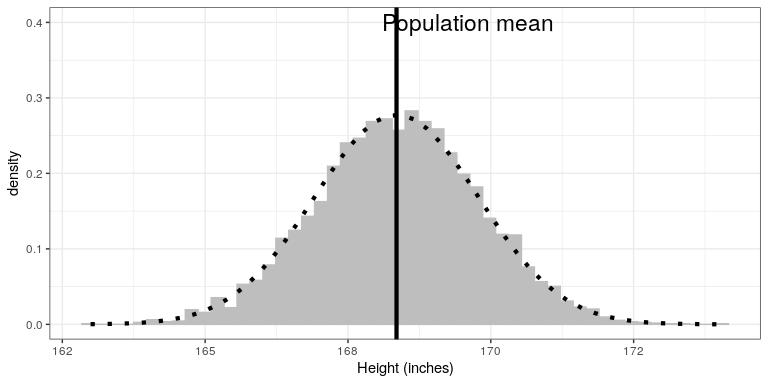
}Ahora vamos a trazar la distribución de muestreo. También se sobrepondremos la distribución muestral de la media predicha sobre la base de la media poblacional y la desviación estándar, para mostrar que describe adecuadamente la distribución muestral real.
# pipe the sampMeans data frame into ggplot
sampMeans %>%
ggplot(aes(meanHeight)) +
# create histogram using density rather than count
geom_histogram(
aes(y = ..density..),
bins = 50,
col = "gray",
fill = "gray"
) +
# add a vertical line for the population mean
geom_vline(xintercept = mean(NHANES_adult$Height),
size=1.5) +
# add a label for the line
annotate(
"text",
x = 169.6,
y = .4,
label = "Population mean",
size=6
) +
# label the x axis
labs(x = "Height (inches)") +
# add normal based on population mean/sd
stat_function(
fun = dnorm, n = sampSize,
args = list(
mean = mean(NHANES_adult$Height),
sd = sd(NHANES_adult$Height)/sqrt(sampSize)
),
size = 1.5,
color = "black",
linetype='dotted'
)